News
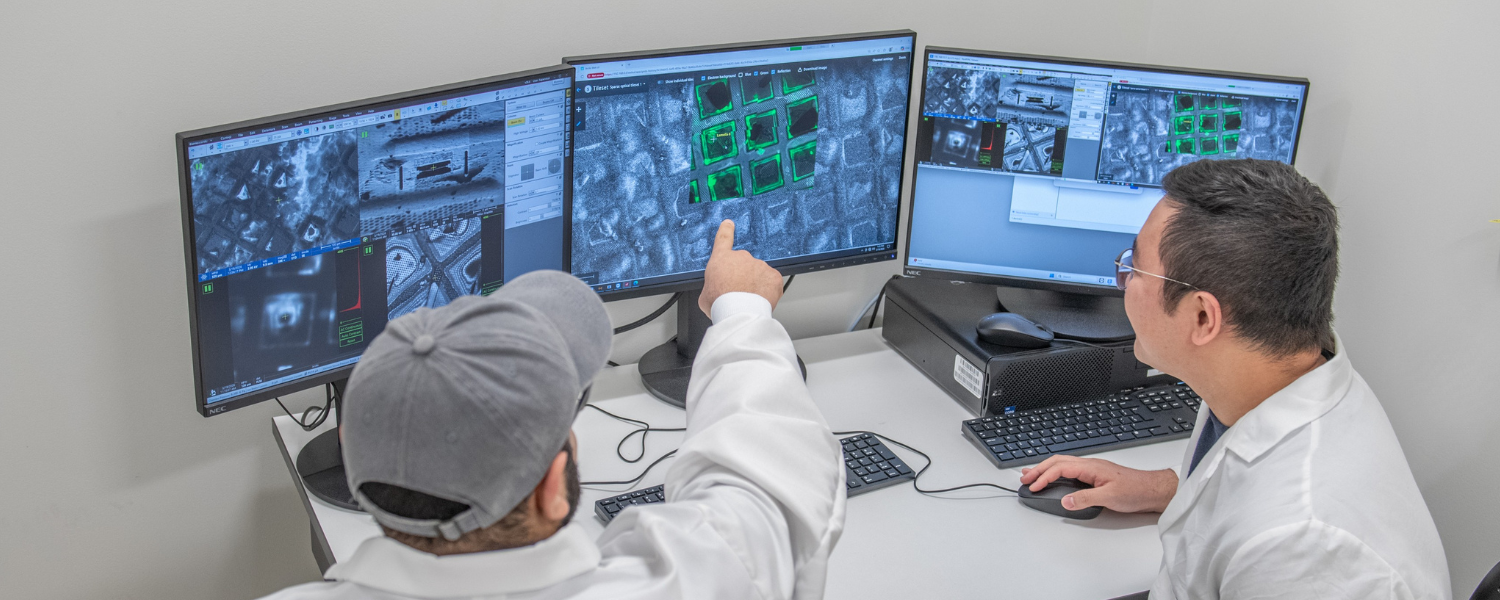

Cutting thinner, seeing deeper: The power of cryo-PFIB

Van Andel Institute and GVSU partner to advance AI-driven biomedical research through newly launched Ph.D. in Computing

How the science of learning is transforming education

How understanding genetics may fuel new ALS therapies

Van Andel Institute and GVSU partner to advance AI-driven biomedical research through newly launched Ph.D. in Computing

Van Andel Institute to recognize Dr. Glenda Halliday with the 2026 Jay Van Andel Award for Outstanding Achievement in Parkinson’s Disease Research

CRISPR variant selectively targets tumor DNA
Van Andel Institute, Cure Parkinson’s renew funding for Parkinson’s clinical trials program
We’re here to help you tell your story.
Van Andel Institute is home to experts in a range of fields, including epigenetics, neurodegeneration, metabolism, structural and cellular biology, and K–12 STEM education.
The Institute’s Communications and Marketing Department assists journalists with information about the groundbreaking research happening at VAI. We can arrange interviews with VAI scientists and provide supporting materials, such as fact sheets, photographs and B-roll video to support the production of news packages. For press inquiries, please reach out to us at [email protected] or submit an inquiry below.
Media contacts
Beth Hinshaw, M.S.
Director, Communications & Marketing
616.234.5519
[email protected]
Zane McMillin
Multimedia Communications Manager, Communications & Marketing
[email protected]
Connect with us

