Near-atomic map of parathyroid hormone complex points toward more targeted therapies for osteoporosis, cancer
April 10, 2019

GRAND RAPIDS, Mich. (April 11, 2019) — An international team of scientists has mapped a molecular complex that could aid in the development of better medications with fewer side effects for osteoporosis and cancer.
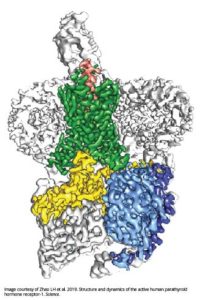
The near-atomic resolution images depict parathyroid hormone receptor-1 (PTH1R), a molecule that conveys signals to and from cells, interacting with two key messengers — a molecule that mimics parathyroid hormone, one of the most important regulators of calcium levels in the body, and a stimulatory G protein, a molecule that mediates bone turnover.
The findings, published today in Science, give researchers a better blueprint for designing drugs for osteoporosis and other conditions such as chachexia, which causes severe weakness and weight loss that can be fatal in cancer patients.
Globally, more than 200 million people have osteoporosis and even more have low bone density. In the coming years, public health experts expect these numbers to rise, fueled in part by an aging population. They also fear an increase in osteoporosis-related fractures due to fewer people taking current medications out of concern over rare side effects.
“The understanding of how all of these molecules fit together has been a missing piece of the puzzle since the discovery of parathyroid hormone 80 years ago,” said H. Eric Xu, Ph.D., a professor at Van Andel Research Institute (VARI) and co-corresponding author of the study. “It’s a big step forward that we hope will one day help people around the world.”
PTH1R is a molecular communication conduit between cells and their environments that fosters development of the bones, skin and cartilage, and regulates levels of calcium in the blood.
To do this, it interacts with molecular messengers such as the parathyroid hormone, which ensures the blood stream has the appropriate amount of calcium to maintain healthy function.
However, too much parathyroid hormone can wreak havoc on the body, spiking the amount of calcium in the blood to dangerous levels, promoting the formation of kidney stones and leaching calcium from bones, which can cause devastating fractures. Too little bogs down metabolism, and contributes to fatigue, weight gain, depression and a host of other issues.
Today’s findings also provide insight into G protein–coupled receptors (GPCRs), a family of signaling molecules to which PTH1R belongs. Taken together, GPCRs are targeted by nearly 30 percent of medications currently on the market.
 GPCRs are notoriously difficult to visualize using traditional X-ray crystallography methods; to date, only about 40 out of more than 800 total GPCRs have had their structures determined. To visualize today’s structure, the team used a groundbreaking technique called cryo-electron microscopy (cryo-EM), which is capable of imaging molecules in unprecedented clarity and can more easily image molecules like GPCRs that are embedded in the cell membrane.
GPCRs are notoriously difficult to visualize using traditional X-ray crystallography methods; to date, only about 40 out of more than 800 total GPCRs have had their structures determined. To visualize today’s structure, the team used a groundbreaking technique called cryo-electron microscopy (cryo-EM), which is capable of imaging molecules in unprecedented clarity and can more easily image molecules like GPCRs that are embedded in the cell membrane.
Xu is co-corresponding author along with Yan Zhang of Zhejiang University School of Medicine, Jean-Pierre Vilardaga of University of Pittsburgh School of Medicine and Ming-Wei Wang of Shanghai Institute of Materia Medica (SIMM) and Fudan University. Xu also holds an appointment at SIMM. Li-Hua Zhao of SIMM, Shanshan Ma of SIMM and University of Chinese Academy of Sciences, Ieva Sutkeviciute of University of Pittsburgh School of Medicine and Dan-Dan Shen of Zhejiang University School of Medicine are co-first authors. Other authors include X. Edward Zhou, Parker de Waal and Karsten Melcher of VARI; Chen-Yao Li, Yanyong Kang, Dehua Yang, Antao Dai, Xiaoqing Cai and Yi Jiang of SIMM; Lisa J. Clark, Frederic G. Jean-Alphonse and Alex D. White of University of Pittsburgh School of Medicine; Jian Chen and Cong Li of Fudan University; and Tomoyuki Watanabe and Thomas J. Gardella of Massachusetts General Hospital and Harvard University.
Research reported in this publication was supported by the Shanghai Science and Technology Development Fund award no. 18ZR1447800 (Zhao) and award no. 16ZR1407100 (Dai); the Young Innovator Association of the Chinese Academy of Sciences award no. 2018325 (Zhao); SA-SBIS Scholarship Program (Zhao); Fudan–SIMM Joint Research Fund award no. FU-SIMM-20174003 (Zhao and Wang); the National Key R&D Program of China award no. 2018YFA0507000 (Wang); the National Natural Science Foundation of China award no. 31300245 (Zhao) and award nos. 81573479 and 81773792 (Yang); Ministry of Science and Technology award nos. 2012ZX09301001, 2012CB910403, 2013CB910600, XDB08020303 and 2013ZX09507001 (Xu); Novo-Nordisk-CAS Research Fund Foundation award no. NNCAS-2017-1-CC (Yang); Van Andel Institute (Xu); National Institute of General Medical Sciences award no. GM127710 (Xu) and T32GM008424 (White); National Institutes of Health award nos. R01DK102495, R01DK111427 and R01DK116780 (Vilardaga); and the Cotswold Foundation Fellowship Awards (Jean-Alphonse and White). The content is solely the responsibility of the authors and does not necessarily represent the official views of the granting organizations.
