
Spatial Collaborative Services
Spatial Collaborative Services (RRID:SCR_027861) combine expertise and technology from across VAI’s Core Technologies and Services to offer investigators a seamless pipeline for spatial imaging.
Spatial technologies bring tissue-specific and subcellular context to gene expression and protein studies, supporting discovery and hypothesis-driven research. The VAI Spatial Collaborative Services group partners with investigators to provide expert experimental design, hands-on technical support and access to state-of-the-art platforms, enabling high-resolution, data-rich insights in an efficient and collaborative environment. For questions, please email [email protected].
Services
- Project consultation and experimental design
- Guidance on targeted panel design (RNA and protein)
- Biobank and biospecimen access
- Tissue processing and embedding
- Microtomy
- Staining (H&E and IF)
- QC services (DV200 and RIN assessment, screening H&E)
- Assay processing and library preparation
- High-resolution imaging
- Data processing and quality control
- Spatial analysis and interpretation support
- End-to-end technical execution and troubleshooting
Equipment
Xenium is a high-resolution, imaging-based spatial profiling system that can characterize up to 5,000 genes and multiplexed proteins in the same tissue section. This system combines spatial fidelity with subcellular resolution in FFPE and fresh frozen tissue, facilitating experiments desiring detailed cell type characterization or exploring cell-cell interactions and transcriptional pathways within their biological contexts. Tissues can be assayed using pre-designed, validated disease or tissue-focused gene panels, fully customized panels, or a mix of the two.
Processing starts with tissue immobilized on a custom Xenium slide, which contains 10.5 x 22.5 mm of surface area to assay multiple tissue sections at a single time; up to two slides can be prepared per run. Probes are hybridized to RNA in the tissue, amplified, and sequenced via cyclic fluorescent images taken at a 200nm per-pixel resolution. Data is visualizable via desktop software from 10X or third parties to localize RNA and protein signals and perform secondary analysis.
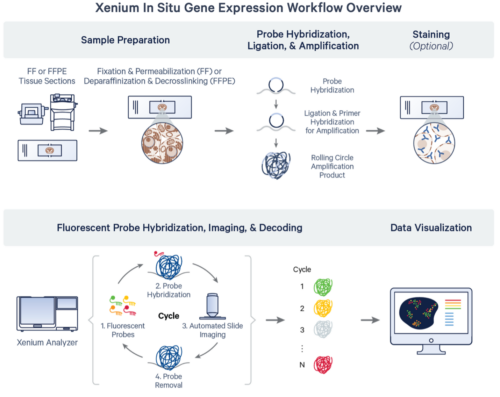
Source: 10x Genomics Xenium In Situ Gene Expression Protocol Planner (CG000601 rev F)
The Visium spatial transcriptomics platform enables whole tissue, spatially resolved, gene expression profiling in both fresh frozen and FFPE material. The system provides transcriptomic information within the context of tissue architecture, tumor microenvironments and cell groups.
The platform relies on custom Visium slides, with either two or four capture areas upon which tissue sections are affixed. Capture areas are either 6.5 x 6.5 mm with 5000 tiled capture spots or 11 x 11 mm with 14,00 capture spots. Each spot has an average capture radius of 50 uM and contains millions of oligonucleotides with a polyA capture sequence, a unique molecule identifier (UMI) and an indexed spatial barcode. After capture, Illumina-sequencing compatible libraries are generated from the probe products and used for high-throughput sequencing.
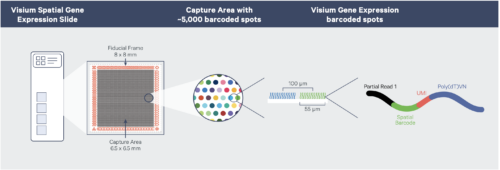
10X Visium v1 workflow (courtesy of 10x Genomics)
The VisiumHD spatial transcriptomics platform expands on the Visium v1 platform to enable whole tissue, spatially resolved gene expression in FFPE material. The system provides higher-resolution transcriptomic information with no dropout areas, allowing for near-single-cell assessment of tissue architecture, microenvironments and co-expression networks.
The assay relies on Visium HD slides containing two capture areas, each 6.5 x 6.5 mm. Capture areas are arrayed with a continuous 2×2 uM lawn of barcoded oligonucleotides. The assay utilizes probe-based gene expression chemistry; probes are hybridized to H&E- or IF-stained FFPE material on a glass slide and imaged to capture spatial orientation. The 10x CytAssist instrument then is used to transfer the transcriptomic probes to the Visium HD slide and extended to add location-specific barcodes. After capture, Illumina sequencing compatible libraries are generated from the probe products and used for high-throughput sequencing.
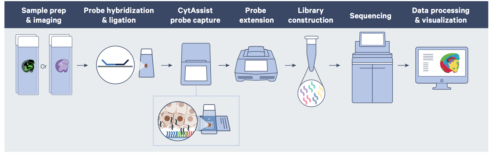
Source: 10x Genomics Grant application Resources for Visium Spatial Gene Expression
The GeoMx DSP enables spatially resolved, region of interest targeted, profiling of both mRNA and proteins in both fresh frozen and FFPE tissue.
The assay is performed on standard negatively charged slides upon with the tissues of interest are mounted; antibodies or RNA probes coupled to photo-cleaveable barcode oligos are incubated with both the tissue and morphology markers. After staining, regions of interest (ROI) can be chosen and optionally subsectioned based on morphology. Oligos from the selected regions are then released by exposure to UV light, incorporated into an Illumina sequencer compatible library and used for high throughput sequencing.
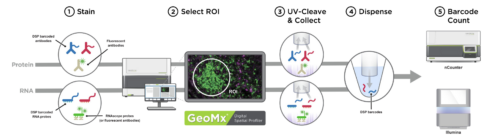
Nanostring GeoMx DSP workflow (Courtesy of Nanostring)
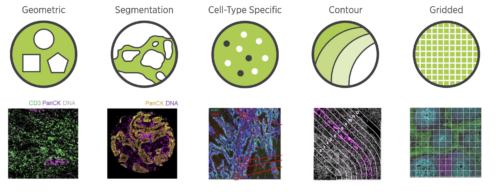
Available Nanostring Region of Interest (ROI) segmentation options (courtesy of Nanostring)
Frequently Asked Questions
Please fill out this form or click the button below and a core member will be in touch to schedule an initial planning meeting with the Spatial group, which includes individuals from the Genomics, Histology, Imaging, and Informatics.
Start Your Project-
- Embedding/re-embedding of fresh frozen/FFPE samples
- Sample quality checks (RIN/DV200 scores, H&E pathology review)
- Sectioning and section placement
- Optimization for permeabilization or antibody times
- Morphology checks (GeoMx only)
- Expression analysis
This includes imaging, slide work, library preparation, sequencing, and basic data analysis.
| Platform | Sample Type | Submission Format | Presubmission QC | Imageable area | Species Compatibility | RNA Compatible | Protein Compatible | Regions Assayed | Resolution | Detection | Deliverables |
| 10x Genomics Visium v1 | Fresh Frozen or FFPE | Blocks or prepared 10x custom slides | RIN/DV200 score: Fresh = RIN >7.0; FFPE = DV200>50% | 6.5 mm x 6.5 mm. 4 regions/ one time use slide | Fresh Frozen = All eukaryotes; FFPE = Human/Mouse | Yes | FFPE only | Whole slide | ~55uM diameter (1-10 cells) | NGS | Fastq files appropriate for Space Ranger or third party pipeline |
| 10X Genomics Visium HD | Fresh Frozen, Fixed Frozen, or FFPE | Blocks or freshly cut charged slides | FFPE = DV200>50% | 6.5 mm x 6.5 mm 2 regions/slide | Human/Mouse | Yes | No | Whole slide | 2x2 uM grid | NGS | Fastq files appropriate for Space Ranger or third party pipeline |
| 10X Genomics Xenium | FFPE | Blocks or prepared 10X custom slides | FFPE = DV200>50% | 10.5 x 22.5 mm/slide | - Predesigned panels: Human/mouse - Custom Panels: species agnostic | Yes | Yes | Whole slide | Subcellular: Transcript XY-localization precision < 30 nm and Z-localization precision < 100 nm Pixel size = ~0.2 µm/pixel | Imaging | Cell Feature matricies, Transcripts (CSV), Tissue Images |
| Nanostring GeoMx DSP | Fresh Frozen, FFPE, Fixed Frozen | Blocks or freshly cut charged slides | None | 35 mm x 14 mm | Human/Mouse | Yes | Yes | Targed regions of interest, user defined | 200-600uM typical (50-100 cells) | Fastq + DSP localization files |
Questions about 10X Visium HD and HD 3’
The following samples can be used in a 10X Visium HD and HD 3’ experiment:
- HD: Fresh frozen, fixed frozen or FFPE samples of human or mouse origin
- HD 3’: Fresh frozen, OCT embedded samples of eukaryotic origin, species agnostic
Visium HD FFPE Tissue Preparation Handbook
FFPE: Please follow standard formalin fixation and embedding protocols. Fresh tissue should be kept in an isotonic solution and fixed within four hours of collection. Following fixation, samples should be stored at to preserve RNA quality. Sections should be <6.5 mm x 6.5 mm to accommodate for the total capture area of one sample. Optimal tissue thickness is 5 µm.
Visium HD Fresh Frozen Tissue Preparation Handbook
Follow the Sample Fixation guidelines in the handbook. Sections should be <6.5 mm x 6.5 mm to accommodate for the total capture area of one sample. Optimal tissue thickness for fresh frozen is 5 µm.
Visium HD Fixed Frozen Tissue Preparation Handbook
Follow the Sample Fixation guidelines in the handbook. Sections should be <6.5 mm x 6.5 mm to accommodate for the total capture area of one sample. Optimal tissue thickness for fresh frozen is 10 µm.
FFPE: RNA quality assessment with DV200 of > 30%. DAPI and H&E staining (nuclei and tissue morphology).
Fresh Frozen: RNA quality assessment with RIN ≥ 4. DAPI and H&E staining (nuclei and tissue morphology).
Fixed Frozen: RNA quality assessment with DV200 ≥ 50%. DAPI and H&E staining (nuclei and tissue morphology).
The barcoded slides contain a 6.5 mm x 6.5 mm continuous grid of 2 µm squares; each 2 µm square contains a unique spatial barcode. Thus, the spatial resolution of the Visium HD assay is 2 µm, which is near single cell resolution.
The Visium HD slides contain two 6.5 mm x 6.5 mm capture areas per slide. Two slides are processed in parallel per run for a total of four capture areas per run (4x reaction kit). There are also 16 reaction kits available, which equates to eight slides or 16 capture areas; however, only two slides may be processed at a time.
Questions about 10X Xenium
FFPE and fresh frozen samples are compatible with both assays.
Please consult the appropriate tissue prep guide for the sample type (see below). Both of the following tissue prep guides are compatible with both Xenium v1 and Xenium prime:
Tissue Prep Guide Xenium v1 Fresh Frozen ➔
Tissue Prep Guide Xenium v1 FFPE ➔
Optimal section thickness is 5 µm for FFPE and 10 µm for fresh frozen samples.
For both Xenium v1 and Xenium Prime, there is one sample area per slide measuring 10.45 mm x 22.45 mm. This sample area can accommodate as many tissues as can fit without obstructing the fiducials.
For both Xenium v1 and Xenium Prime:
DAPI and H&E staining of sections from both FFPE and fresh frozen sample types to assess tissue and nuclei morphology.
RNA quality should be assessed with a threshold DV200 score of >30%.
The spatial resolution of both Xenium v1 and Xenium Prime is subcellular (30 nm). The Xenium assays are our highest spatial resolution assays at the tradeoff of assaying fewer genes than Visium.
The Xenium v1 and Xenium Prime slides are equivalent slides that contain one capture area per slide with a sample area measuring 10.45 mm x 22.45 mm. As the reagent kits are 2x reaction kits, the Xenium v1 assay is always performed with two Xenium slides in unison.
Question about Nanostring GeoMx DSP
The following samples can be used in a Nanostring GeoMx DSP experiment: FF or FFPE, OCT from mouse or human tissues only.
-
- Fresh frozen samples: Flash freeze in liquid nitrogen cooled isopentane as soon as possible after collection. Optimally embed in OCT or CMC either at the same time or afterward.
- FFPE samples: Please follow standard formalin fixation and embedding protocols. Samples should ideally be stored at 4°C post fixation to preserve RNA quality.
Optimal tissue thickness is 5um.
Tissue should be sectioned onto positively charged slides no more than two weeks prior to running the Nanostring DSP assay.
Morphology check: A test slide for a scanning only run (without ROI definition, UV illumination, or ROI collection) is strongly recommended to expedite ROI selection the day of the experiment and validate the fluorescent staining of your morphology markers. All requests utilizing custom antibodies are required to undergo this process.
Minimum of 50–100 cells per ROI (region of interest). May be subdivided into multiple AOIs (areas of interest) based upon morphology markers in that region.
Multiple sections or sections from multiple tissues can be placed on the same slide, so long as the same probe set will be used on all slides.
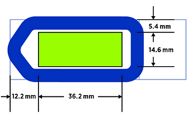
All sections must fit within the 40mm x 17mm slide region and be entirely within the GeoMx gasket to ensure a good seal.
-
- Leica Bond Plus (Cat# S21.2113.A)
- Fisherbrand Superfrost Plus (Cat # 12-550-15)
-
- Whole transcriptome atlas (human or mouse)
- Cancer transcriptome atlas (human)
- Protein assays with NGS readout (human or mouse)
Team
Spatial Collaborative Services comprises experts across VAI Core Technologies and Services. Additional support is provided by the VAI Pathology and Biorepository Core through its Histology and Molecular Processing team.

Corinne Esquibel, Ph.D.
Director, Optical Imaging Core

Marie Adams, M.S.
Director, Genomics Core

Lisa Turner, B.S., HT, QIHC, QBRS, QLS (ASCP)
Core Associate, Pathology and Biorepository Core

Katelyn Becker, M.S.
Core Laboratory Manager, Genomics Core

Michael Davis
Core Associate, Genomics Core

Lorna Cohen, Ph.D.
Core Laboratory Manager, Optical Imaging Core

Jian Wei Tay, B.Sc. (Hons I), Ph.D.
Core Bioinformatics Scientist II, Optical Imaging Core

Daisy Fu, Ph.D.
Core Bioinformatics Scientist II, Bioinformatics and Biostatistics Core

