Research brief: Four molecular subtypes revealed in rare bile duct cancer
March 13, 2017
The short version
Scientists now have a new resource for studying a rare bile duct cancer that may serve as a blueprint for placing patients in the appropriate clinical trials.
Why are these findings important?
Properly dividing patients into appropriate trial groups based on disease subtype is critical for accurately studying the safety and efficacy of potential new therapies.
Clinically, a detailed understanding of disease subtypes may one day help physicians decide which treatments are best for individual patients and avoid approaches that may not work.
Background
Cholangiocarcinoma is a rare bile duct cancer that affects 8,000 people a year in the U.S.—a number that has been rising in recent years—and is related to conditions such as hepatitis B or C that cause inflammation of the liver or the nearby ducts. Incidence rates are higher elsewhere in the world, particularly in Southeast Asia, where many of the cases are related to infection with parasitic liver flukes. Currently, treatment options are limited and patient prognosis is generally poor.
 How they did it
How they did it
The results came from The Cancer Genome Atlas (TCGA) Research Network, a National Cancer Institute-led multi-institutional effort to define the molecular basis of cancer. This large-scale project leverages the resources of hundreds of experts at institutions across the country to characterize cancers across multiple platforms, including genetic mutations and epigenetic aberrations.
The TCGA Research Network studied 38 samples, 89 percent of which were derived from North America and 97 percent of which were negative for hepatitis B and C. None of the samples were related to parasitic infection.
The team analyzed the samples across five platforms—or types of tests—including whole exome sequencing, SNP arrays, RNA-sequencing (including microRNA and long noncoding RNA), DNA methylation and reverse-phase protein arrays.
What they found (the long version)
The team identified four molecular subtypes that may aid scientists in studying the disease and that one day may have clinical implications. However, the authors caution that a broader sample size and confirmation in disease models are needed before these subtypes are used by physicians in patient treatment.
One of the subtypes featured mutations in the IDH1 and IDH2 oncogenes, resulting in the hypermethylation and subsequent silencing of another gene called ARID1A, which plays a role in transcription regulation by chromatin remodeling. Mutations in IDH also have previously been linked to dysfunction in cellular metabolism, thus providing another avenue for therapeutic development.
Interestingly, the team also found a group of liver tumors that not only had a similar molecular profile to cholangiocarcinoma but that also had IDH mutations, supporting the view that liver tumors fall on a spectrum. Given its similarity to cholangiocarcinoma, the authors recommend further study into this group of liver tumors to ensure accurate classification.
Overall, the findings show cholangiocarcinoma to be incredibly diverse in terms of genetic and epigenetic alterations, indicating that it is unlikely that treatment with a single drug will lead to sustained response or remission. More likely, a combination of drugs based on an individual’s specific tumor will be needed.
Scientist’s take
“Unlike cancers that are largely marked by one type of genetic or epigenetic alteration, cholangiocarcinomas tend to involve a collection of errors across multiple important pathways. To complicate things further, geographic location, presence of parasitic infection or hepatitis and location in the ducts also play critical roles,” says Peter W. Laird, Ph.D., a professor at Van Andel Research Institute and member of the TCGA Research Network. Dr. Laird led the epigenetic analysis for the study. “It is our hope that this paper will serve as a reference for our colleagues who are working on potential new therapies for this disease.”
“The collaborative nature of this project allowed us to look at cholangiocarcinoma across multiple platforms, a feat that is not always possible alone,” says Toshinori Hinoue, Ph.D., a bioinformatics scientist in the Laird Laboratory. “As a resource, this paper will provide clarity for future studies.”
Learn more
The paper was published today in Cell Reports. You can find it here. Read more from TCGA here.
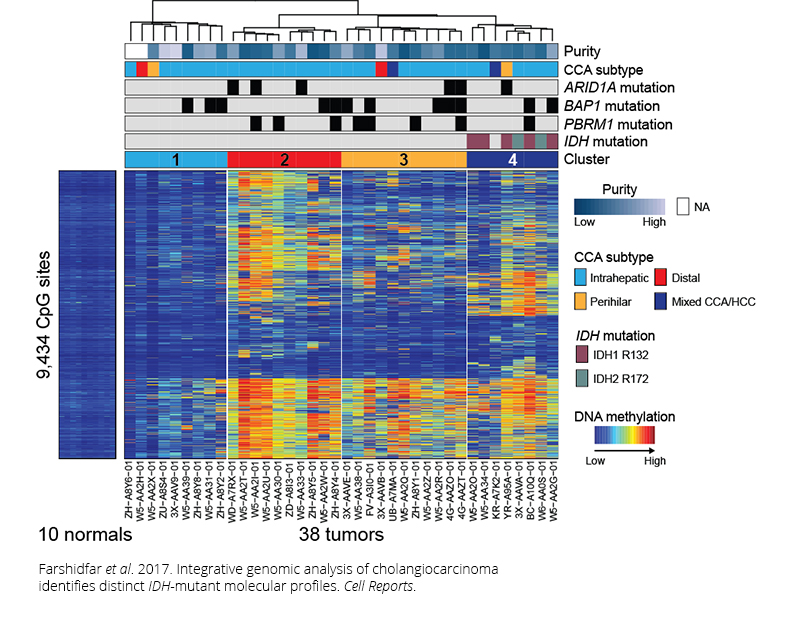
Farshidfar F et al. 2017. Integrative genomic analysis of cholangiocarcinoma identifies distinct IDH-mutant molecular profiles. Cell Rep.
KEY TERMS
Bile ducts: The bile ducts are tubes that facilitate the movement of bile from the liver to the small intestine, where it aids in digestion.
Epigenetics: Literally meaning “on top of genetics,” epigenetics refers to the factors that help control gene expression. You can find a good crash course in epigenetics here.
Oncogenes: Genes that are typically associated with cancer
DNA Methylation: An epigenetic mechanism in which molecules called methyl groups are added to DNA, ultimately determining whether a gene—a unit of DNA with specific instructions for a function—is active or silent. Hypermethylation, meaning a surplus of methyl groups have been added to a gene, often are present in cancer. Learn more about DNA methylation here.
Transcription: The first step in the process of gene expression, in which genetic information flows from DNA to RNA.
