Cryo-EM reveals interaction between major drug targets
Structure provides “blueprint” for drug development, answers a longstanding question in biology
June 12, 2018
GRAND RAPIDS, Mich. (June 13, 2018) — For the first time, scientists have visualized the interaction between two critical components of the body’s vast cellular communication network, a discovery that could lead to more effective medications with fewer side effects for conditions ranging from migraine to cancer.
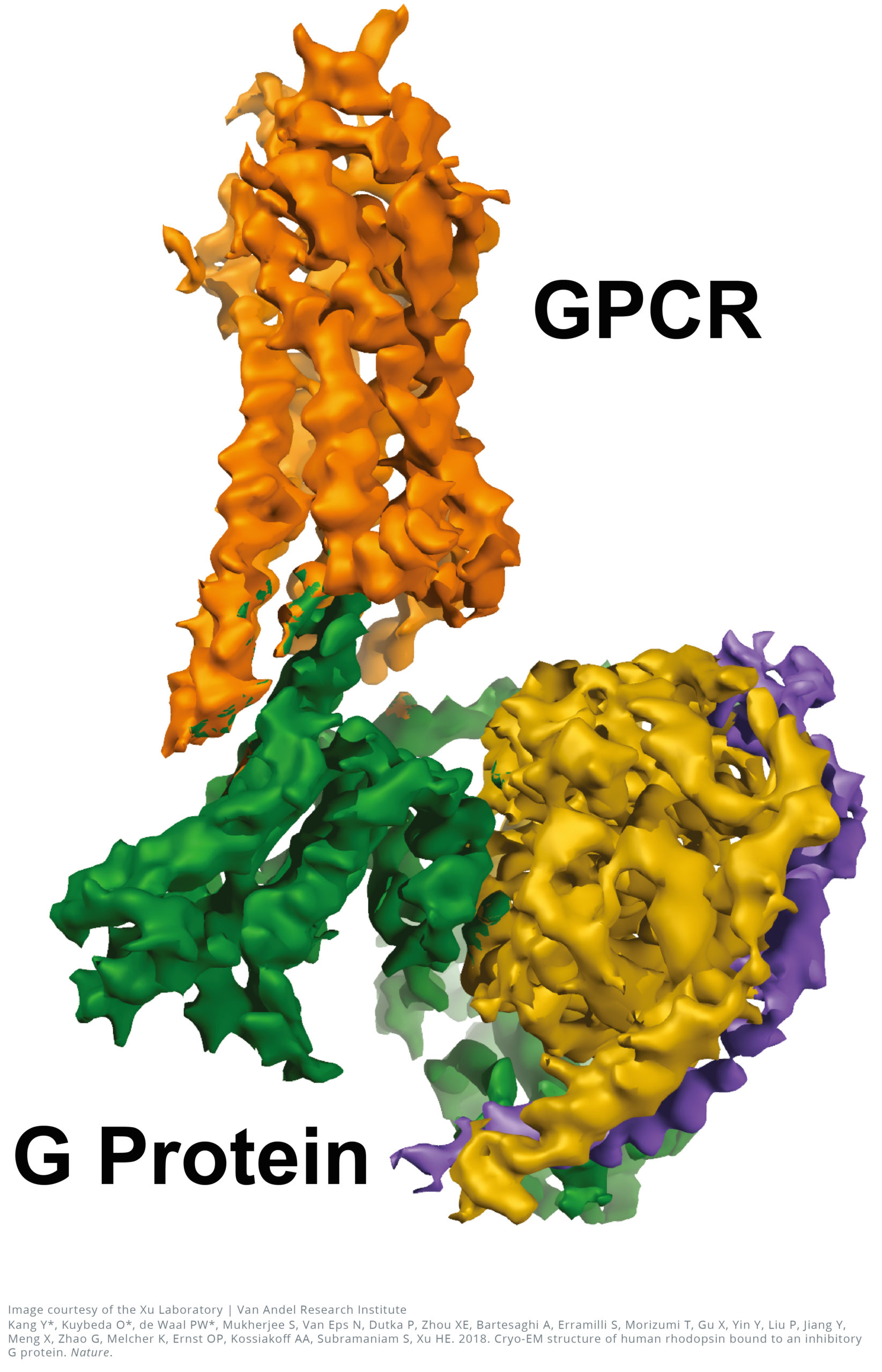
The near-atomic resolution images, published today in Nature, show a G-protein coupled receptor (GPCR) called rhodopsin bound to an inhibitory G protein, and provides a blueprint for designing more precise, selective drugs while also solving a longstanding problem in the field.
“Visualizing this complex resolves a missing chapter in the GPCR story by finally revealing how these two molecules interact in exquisite detail,” said H. Eric Xu, Ph.D., a professor at Van Andel Research Institute (VARI) and one of the study’s senior authors. “Everything in biology is based on molecular interactions so the more we know about how the structures of these two molecules work together, the better position we are in to design improved medications with fewer undesired effects.”
Today’s findings were made possible through use of a revolutionary technique called cryo-electron microscopy (cryo-EM), which allows scientists to see tough-to-visualize molecules in startling clarity.

“The use of cryo-EM technology to obtain structural information on important pharmaceutical targets such as GPCRs in various states demonstrates that we are now in a position to apply these methods for drug discovery applications,” said Sriram Subramaniam, Ph.D., an investigator at the National Cancer Institute of the National Institutes of Health and a senior author of the study.
Embedded in the cell membrane, GPCRs act as conduits between a cell and its environment, interacting with G proteins and other signaling molecules called arrestins to convey important messages to and from the cell that regulate a gamut of physiological functions, including growth, immune responses and sensory perception.
When linked up with GPCRs, inhibitory G proteins regulate the production of secondary chemical messengers that have effects throughout the body, from interactions with serotonin receptors in the brain and gut, which help regulate mood and appetite, to interactions with dopamine receptors in the brain, which control reward responses and voluntary movement, among many others.
These wide-ranging interactions with G proteins and arrestins, coupled with their accessibility on the outside of the cell, make GPCRs attractive targets for therapeutic development. Currently, more than 30 percent of medications on the market work by interacting with GPCRs.
“The information revealed by our findings will help facilitate the design of a new generation of medications,” said Yanyong Kang, Ph.D., a research scientist in the Xu Laboratory and co-first author of the study. “Because this is the first GPCR-inhibitory G protein complex to be structurally determined, we believe our methods will help lead to the characterization of other important, yet difficult-to-visualize GPCRs.”
The 3D images generated by the team reveal a specialized helix at the end of the inhibitory G protein that acts as a structural signature, which helps GPCRs like rhodopsin differentiate between inhibitory G proteins and another type of G protein known as a stimulatory G protein.
Today’s findings are the latest in a series of firsts for Xu and his team, which include a landmark 2015 Nature study that first described the structure of rhodopsin and arrestin in complex together. This work, which was hailed as a major breakthrough in the field, earned Xu the Hans Neurath Award from The Protein Society and the Hans Neurath Foundation in 2016.
In a follow-up study published in Cell in 2017, Xu and his collaborators further refined their earlier structure of the rhodopsin-arrestin complex, and revealed a set of phosphorylation codes that dictate the assembly of GPCR-arrestin complexes.
GPCRs are notoriously difficult to visualize using traditional X-ray crystallography methods; to date, only 40 out of more than 800 total GPCRs have had their structures determined, including Xu’s rhodopsin-arrestin complex.
To determine today’s structure, the team harnessed VARI’s high-powered Titan Krios cryo-electron microscope, which is capable of imaging molecules 1/10,000th the width of a human hair and can more easily visualize molecules like GPCRs that are embedded in the cell membrane. The Institute’s Krios, which is part of its David Van Andel Advanced Cryo-Electron Microscopy Suite, is one of fewer than 120 such microscopes in the world.
Subramaniam and his team have pioneered the use of cryo-EM to determine some of the highest resolution structures reported so far using cryo-EM, including several clinically relevant ligand-protein complexes.
This study was a collaboration between teams from VARI, the National Cancer Institute, University of Chicago, University of Toronto and Shanghai Institute of Material Medica.
In addition to Xu, Subramaniam and Kang, authors include Parker de Waal, W. Edward Zhou, Ph.D., Xin Gu, M.S., Yanting Yin, Xing Meng, Ph.D., Gongpu Zhao, Ph.D., and Karsten Melcher, Ph.D., of VARI; Oleg Kuybeda, Ph.D., and Alberto Bartesaghi, Ph.D., of the Cancer Research Technology Program at Frederick National Laboratory for Cancer Research and the National Cancer Institute; Somnath Mukherjee, Ph.D., Prezemyslaw Dutka, M.Sc., Satchal Erramilli, Ph.D., and Anthony Kossiakoff, Ph.D., of University of Chicago; Ned Van Eps, Ph.D., Takefumi Morizumi, Ph.D., and Oliver P. Ernst, Ph.D., of University of Toronto; and Ping Liu and Yi Jiang, Ph.D., of VARI-Shanghai Institute of Materia Medica.
Kang, Kuybeda and de Waal are co-first authors of the study. Xu also is affiliated with VARI-SIMM; Dutka also is affiliated with Jagiellonian University; and Liu also is affiliated with University of the Chinese Academy of Sciences.
Research reported in this publication was supported by the National Institutes of Health under grant number DK071662 (Xu); the American Asthma Foundation (Xu); Van Andel Institute (Xu, Melcher); the Ministry of Science and Technology [China]; National Institute of General Medical Sciences grant numbers GM117372 (Kossiakoff) and GM0875119 (Kossiakoff); Pfizer (Kossiakoff); National Natural Science Foundation grant 31770796 (Jiang); the Canada Excellence Research Chairs Program (Ernst); the Canadian Institute for Advanced Research, the Anne and Max Tanenbaum Chair in Neuroscience (Ernst); Center for Cancer Research, National Cancer Institute (Subramaniam); and Frederick National Laboratory for Cancer Research, National Institutes of Health under contract HHSN261200800001E (Subramaniam). The content is solely the responsibility of the authors and does not necessarily represent the official views of the funding organizations.
